|
If the linking information spans large distances on the chromosome, the resulting scaffolds can span entire chromosomes or chromosome arms. 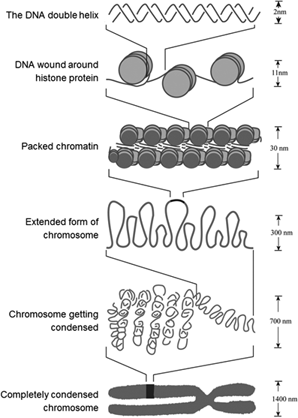
Scaffolding utilizes long-range linking information such as BAC or fosmid clones, optical maps, linked reads, or chromosomal conformation capture to order and orient contigs. Ambiguous reconstructions are output as edges linking unitigs. Unambiguous reconstructions of the sequence are output as “unitigs” (or often “contigs”). Repetitive sequences exceeding the sequencing read length introduce ambiguity and prevent complete reconstruction. Most approaches are based on either a de Bruijn or a string graph formulation. Reads originating from the same locus are identified using assembly software, which can use these overlaps to reconstruct the genome sequence. 
Most genome projects rely on whole genome shotgun sequencing which yields an oversampling of each genomic locus. Genome assembly is the process of reconstructing a complete genome sequence from significantly shorter sequencing reads. This is a PLOS Computational Biology Methods paper. Anthony Schmitt and Siddarth Selvaraj are employees of Arima Genomics, a company commercializing Hi-C DNA sequencing technologies. This work utilized the computational resources of the NIH HPC Biowulf cluster ( ).Ĭompeting interests: Sergey Koren has received travel and accommodation expenses to speak at Oxford Nanopore Technologies conferences. AR was also supported by a grant from the Korean Visiting Scientist Training Award (KVSTA) through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health \& Welfare, Republic of Korea (grant number: HI17C2098). SK, AR, BPW, and AMP were supported by the Intramural Research Program of the National Human Genome Research Institute, National Institutes of Health. JG and MP were supported by NIH grant R01-AI-100947 to MP. AS and SS were funded by generous support from NHGRI (grant\# 1R44HG009584). The work is made available under the Creative Commons CC0 public domain dedication.ĭata Availability: All relevant data are within the manuscript and its Supporting Information files.įunding: The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. This is an open access article, free of all copyright, and may be freely reproduced, distributed, transmitted, modified, built upon, or otherwise used by anyone for any lawful purpose. Received: JanuAccepted: JPublished: August 21, 2019 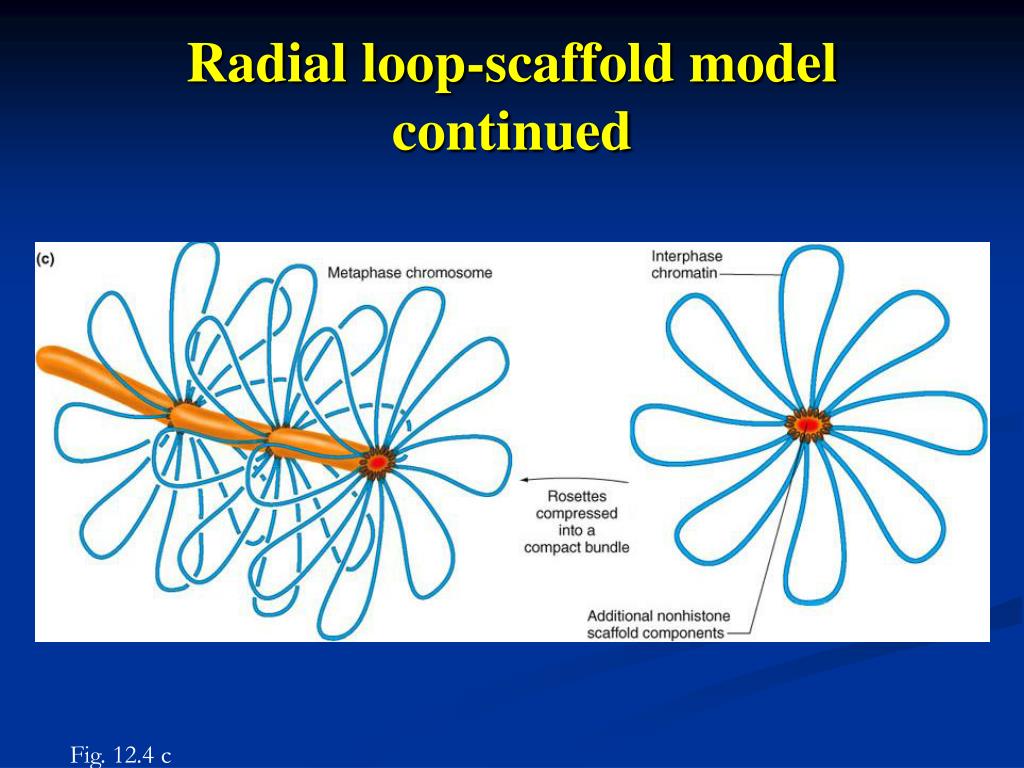
(2019) Integrating Hi-C links with assembly graphs for chromosome-scale assembly. Citation: Ghurye J, Rhie A, Walenz BP, Schmitt A, Selvaraj S, Pop M, et al.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
